
Back in 2017, I was asked to peer review an article and its author asked if I would like the review to be “open” – that is that my name would be shown as a reviewer; [cite]10.1073/pnas.1709586114[/cite/] indeed it was!

Back in 2017, I was asked to peer review an article and its author asked if I would like the review to be “open” – that is that my name would be shown as a reviewer; [cite]10.1073/pnas.1709586114[/cite/] indeed it was!
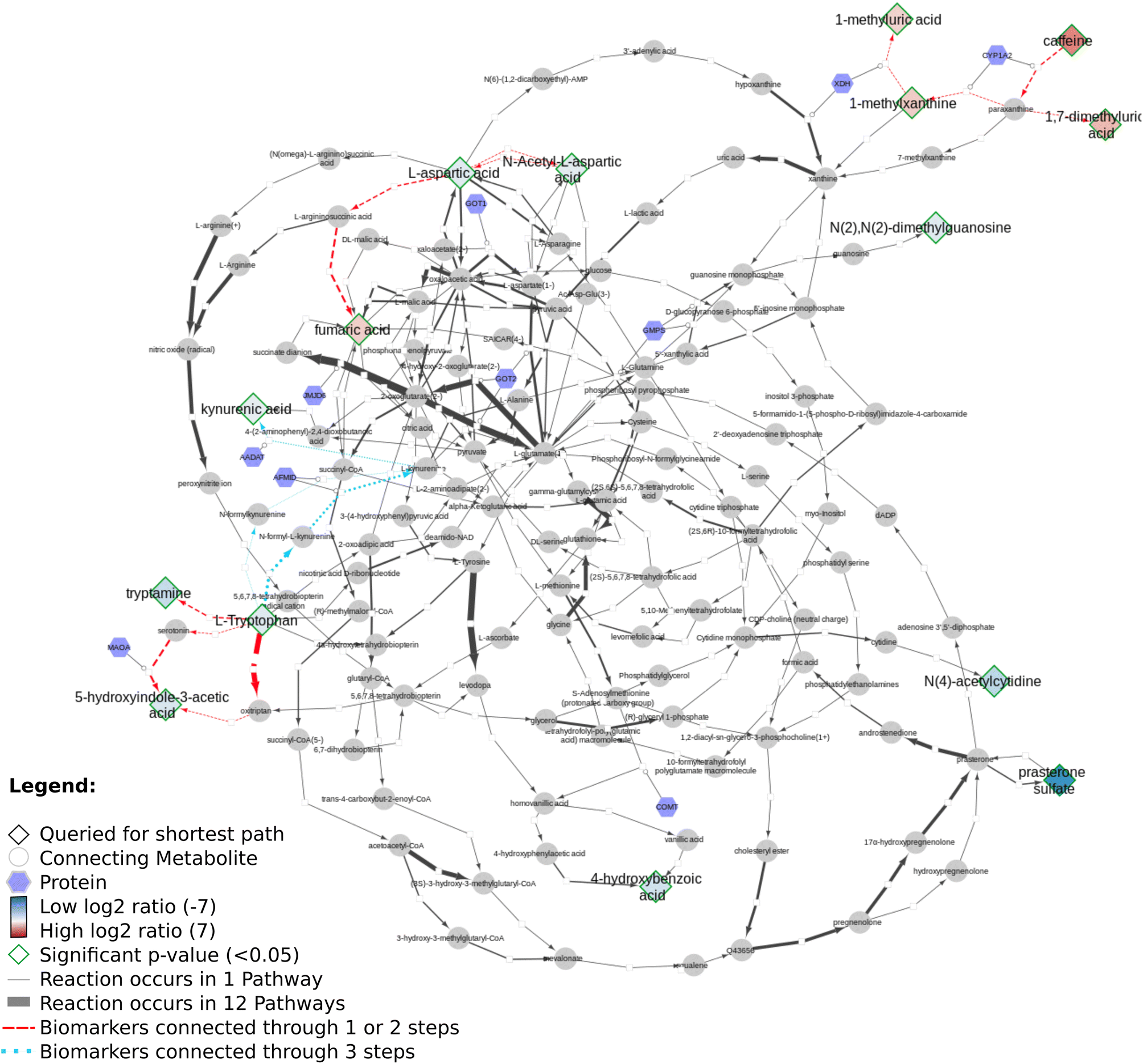
I am still catching up with a lot of work, and found out I actually had forgotten to blog about this cool article by Denise Slenter: “Discovering life’s directed metabolic (sub)paths to interpret human biochemical markers using the DSMN tool” (doi:10.1039/D3DD00069A). This paper explains how various open science resources (Wikidata, Reactome, WikiPathways) are used to visualize the biological story of the data from two metabolomics experiments

About a year ago I started migrating my blogger.com blog to a git-version-controlled, Markdown-based blogging platform. I have to say, it has been a happy year. It actually is awesome to port old blog posts (follow that here) and to see what I have been working on some 17, 18 years ago. I do have a nasty bug to fix that causes the conversion of the Markdown to HTML is scaling badly.

Metadata is something that goes on behind the scenes and is rarely of concern to either author or readers of scientific articles. Here I tell a story where it has rather greater exposure. For journals in science and chemistry, each article published has a corresponding metadata record, associated with the persistent identifier of the article and known to most as its DOI.
I've been playing around with generating non-equilibrium conformations by molecular dynamics recently, and I've been thinking about how to best parse the outputs of a dynamics simulation.

In the CDK2024 grant we wrote about updating various software projects using the Chemistry Development Kit. We even wrote that “[r]equired API changes will be publicly shared and disseminated with the Groovy Cheminformatics with the Chemistry Development Kit book (egonw.github.io/cdkbook/)”. The Groovy Cheminformatics with the Chemistry Development Kit book is a project that has run since 2009.
I should start by saying that the server on which this blog is posted was set up in June 1993. Although the physical object has been replaced a few times, and had been “virtualised” about 15 years ago, a small number of the underlying software base components may well date way back, perhaps even to 1993.
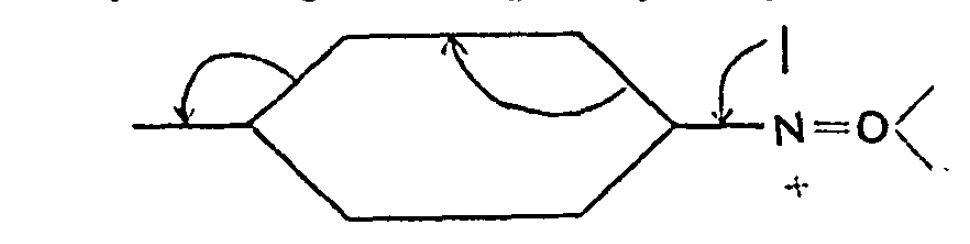
Chemists now use the term “curly arrows” as a language to describe the electronic rearrangements that occur when a (predominately organic) molecule transforms to another – the so called chemical reaction. It is also used to infer, via valence bond or resonance theory, what the mechanistic implications of that reaction are.

Scientific software can be confusing, particularly when you're doing something that the software isn't primarily intended for. I often find myself wanting to run quick-and-dirty molecular dynamics simulations on small organic molecules, but I've struggled to find an easy way to do this using open-source tools.
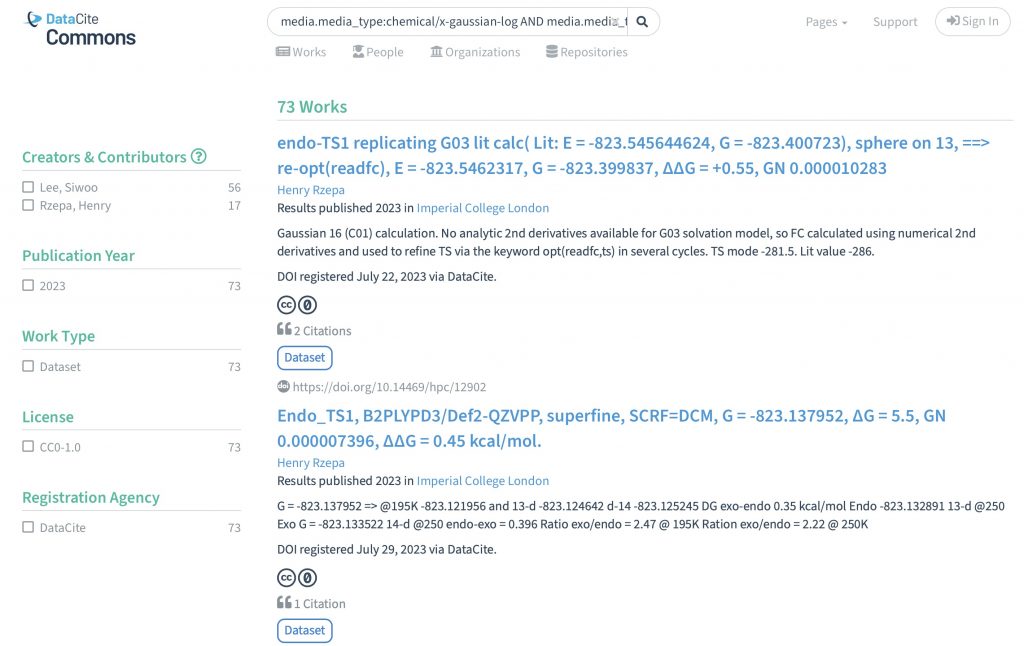
I can remember a time when journal articles carried selected data within their body as e.g. Tables, Figures or Experimental procedures, with the rest consigned to a box of paper deposited (for UK journals) at the British library. Then came ESI or electronic supporting information.