
I've added Index Fungorum to the list of RSS feeds that I generate at bioguid.info/rss.

I've added Index Fungorum to the list of RSS feeds that I generate at bioguid.info/rss.
Although I'd been thinking of getting the wiki project ready for e-Biosphere '09 as a challenge entry, lately I've been playing with RSS has a complementary, but quicker way to achieve some simple integration. I've been playing with RSS on and off for a while, but what reignited my interest was the swine flu timemap I made last week. The neatest thing about the timemap was how easy it was to make.

Well, not Darwin himself, exactly. The Evolution Directory (better known as "EvolDir") is a mailing list run by Brian Golding at McMaster University, Ontario.

ZooKeys (ISSN 1313-2970) is a new journal for the rapid publication of taxonomic names, rather like Zootaxa . On first glance it has some nice features, such as being Open Access (using the Creative Commons Attribution license), DOIs, and RSS feeds -- although these don't validate, partly due to an error at the bottom of the feeds: Warning : Cannot modify header information - headers already sent by (output started at
More followup on feeds. I’ve set up a friendfeed which is available here. Amongst other things I’ve been trying to aggregate my comments on other people’s blogs. The way I am doing this is creating a tag in Google Reader which I have made public. When I leave a comment I try to subscribe to a comment feed for that Blog, I then tag it as ‘Comments feed’ which aggregates and makes it public as an RSS feed.
Following on from (but unrelated to) my post last week about feed tools we have two posts, one from Deepak Singh, and one from Neil Saunders, both talking about ‘friend feeds’ or ‘lifestreams’. The idea here is of aggregating all the content you are generating (or is being generated about you?) into one place. There are a couple of these about but the main ones seem to be Friendfeed and Profiliac.
David Strumfels posted news about the Useful Chemistry CMLRSS feed . He explains how this feed can be accessed using Jmol and Bioclipse. The latter are accompanied by two AVI movies: one about creating a new OPML file, and one about accessing the CMLRSS file from the OPML.
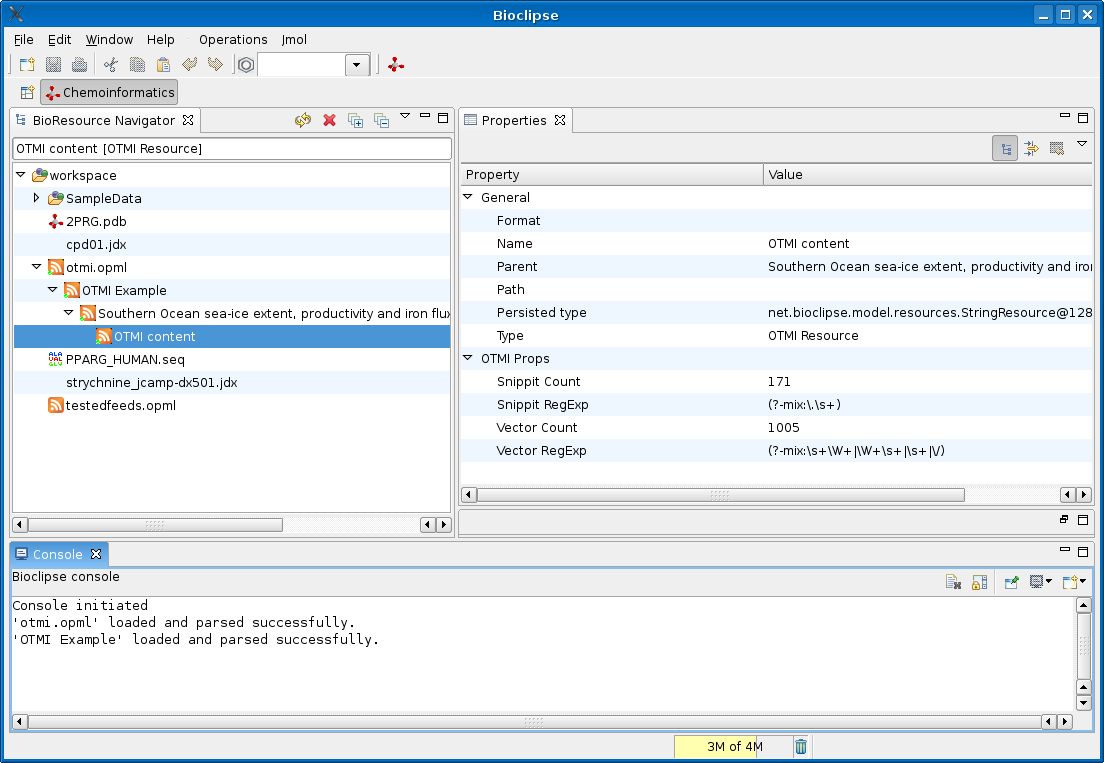
Timo Hannay blogged in Nature’s Nascent blog about the Open Text Mining Interface (OTMI), which is “a suggestion from Nature about how we might achieve text-mining and indexing purposes”. The idea is that each article has a link pointing to a machine readable file containing raw data about (and from?) the article.