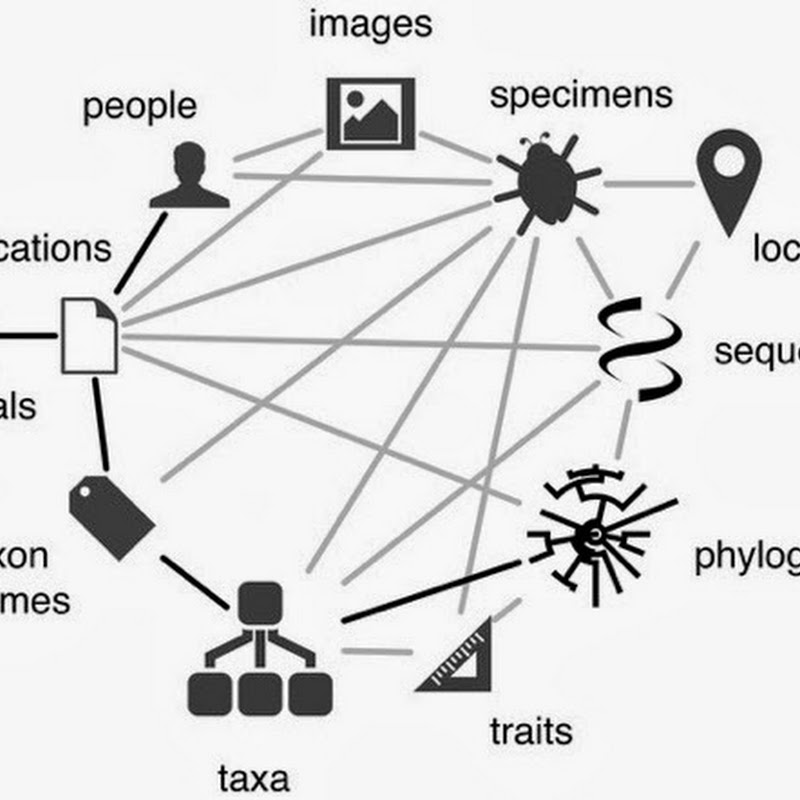
The goal of my BioNames project is to link every taxonomic name to its original description (initially focussing on animal names). The rationale is that taxonomy is based on evidence, and yet most of this evidence is buried in a non-digitised and/or hard to find literature. Surfacing this information not only makes taxonomic evidence accessible (see Surfacing the deep data of taxonomy), it also surfaces a lot of basic biological information.