More research/literature mapping tools - Connected Papers and CoCites

It's a cliche to say that the volume of scientific literature produced is increasing at a rate beyond which researchers can keep up. With the rise of Machine learning techniques and more importantly the increasing availability of open meta-data and increased open full text , we are starting to see tools appear that try to leverage this to produce applications that can help take the load off researchers on what to read and to see connections between papers in a field.
In this blog post, I am going to review 2 new ones that have come across my radar - CoCites (direct link to my review) and Connected Papers (direct link to my review).
CoCites and Connected Papers join the other tools I am familiar with including Bibliometric/Science mapping tools (VOSviewer and CiteSpace), Open Knowledge Maps, Citation Gecko and Local Citation Network. Notice many of these tools use open data sources like Microsoft Academic Graph, Crossref , PubMed or Semantic Scholar Open Research Corpus
I will do a brief summary of these tools and how the two new ones CoCites and Connected Papers are similar or differ from them as well as a brief overview of yet another new one PaperGraph.
You can jump directly to the review of Cocites here and Connected Papers here.
Brief comparison of the tools
NameData SourceInputNetwork graph generatedSuggested papersComment CiteSpace WOS, Scopus, Crossref, Lens.org, and more with conversion Input files Multiple methods (e.g. bibliometric coupling, cocitation, citation) NA More of mapping tool than literature review support VOSviewer Accepts WOS, Scopus,Dimensions, Microsoft Academic, Crossref, COCI, OCC, Wikidata and more Input files or keyword search of Crossref API, Microsoft Academic, Crossref, etc Multiple methods (e.g. bibliometric coupling, cocitation, citation) NA More of mapping tool than literature review support Open Knowledge Maps Base or PubMed Keyword
Co-word graph based on title, journal name, author names, subject keywords, abstract. NA Not citation based Whocites (source) Google Scholar Series of Keyword Records top 10 papers or book from GS search, Using Google Scholar's 'search within citations' it checks to see if any of the authors recorded to the database have cited any of the publications.
NA very slow Citation Gecko Crossref Multiple papers Papers cited by seed papers or citing seed papers Most cited or citing papers by seed papers.
You can iterate and grow network by adding these as seed papers Location Citation Network Microsoft Academic or Crossref One paper or multiple papers References of input paper Most cited by local network Also has co-authorship network CoCites PubMed One paper or multiple papers No graph yet. Uses cocitations of last 100 cited papers of input paper. Sort by co-citations or similarity (% of cocitations) Future version will allow multiple papers Connected Papers Semantic Scholar Open Research Corpus One paper Similarity metric based on concepts of cocitations and bibliometric coupling Additional function to detect "Prior works" (most cited by local network) and "derivative works" (works that cite most of local network Papergraph Semantic Scholar Open Research Corpus One paper 20 X 20 citations or references of input paper- Most cited by local network
A brief listing of existing tools
Why did I list the tools above when they are so many others?
As of time of writing, I'm currently familiar with the following set of tools that fits the following critera.
Has a user friendly interface
Uses an index that is cross-disciplinary (typically Microsoft Academic, Crossref, Scopus etc) or are focused on life sciences (PubMed, PMC) to produce a network visualization of papers. (So no narrow subject specific tool)
Accepts input as keyword or individual/series of papers to generate a network graph of papers and/or recommends papers
I also favour tools that are opensource or at least currently free to use with no subscription version (e.g. Iris.AI).
1. Bibliometric science mapping tools -there's a big bunch of these (e,g. CitNetExplorer , CRExplorer) of which I have tried the most are VOSviewer and CiteSpace - These are tools designed and use by bibliometrician that allow you to map out connections between sets of papers. These tools tend to be complicated to use , with many options (with VOSviewer perhaps the cleanest) as they are geared more for bibliometric research rather than generic literature review support - a typical use is to export a large set of papers using keywords from a database like Scopus or Web of Science and to import the papers into these tools for mapping connections between the papers to get an overview of the field.
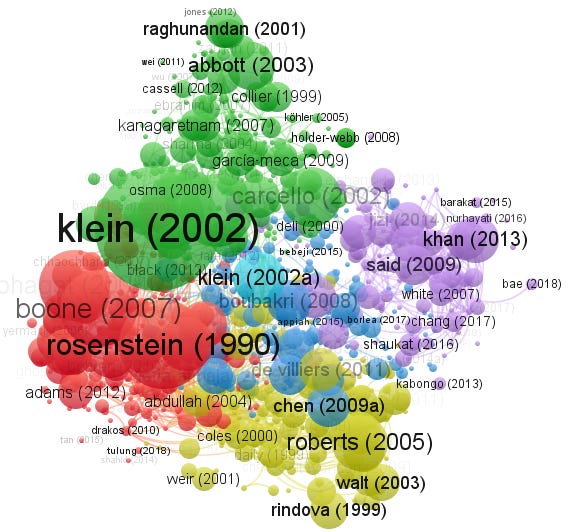
Bibliometric coupling of Dimensions (title and abstract) data exported with Board independence in VOSviewer
VOSviewer impressively supports a large number of indexes besides the usual Scopus/Web of Science it includes Dimensions, Crossref, Microsoft Academic, COCI, OCC, Semantic Scholar and more. It has nice sensible defaults and is easy to use.
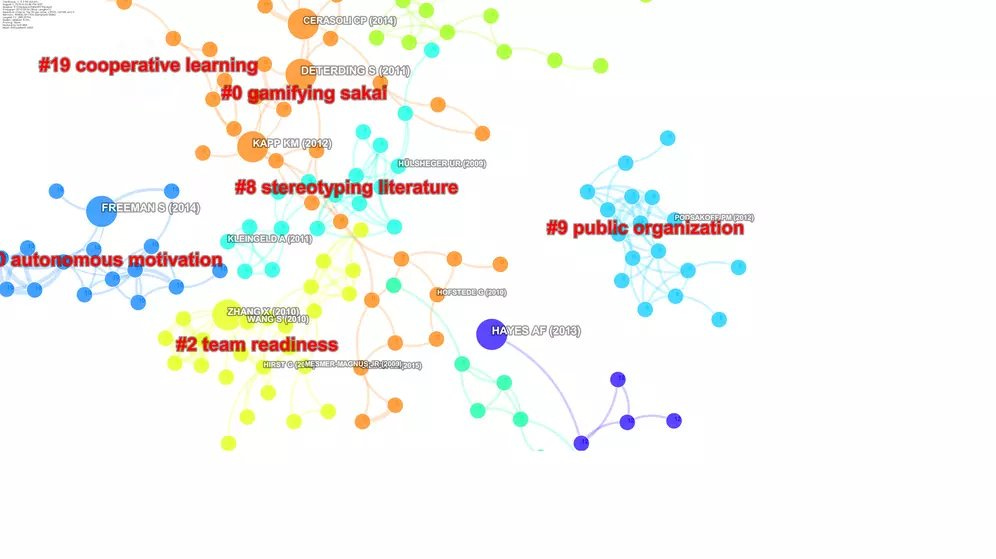
CiteSpace analysis of papers from Scopus on Team Creativity
CiteSpace also supports a large number of indexes, though you have to do more work to convert them. It has a ton of features (at the cost of complexity to use), I personally like the feature of auto labelling of cluster seen above.
2. Open Knowledge Maps (reviewed here) - This is perhaps one of the first of the current generation of tools geared to specifically support researchers with literature review. The current version (my blog post reviews a beta version) of it is somewhat simple, you enter a keyword and it searches either PubMed or BASE and tries to cluster papers based on text similarity of meta-data- title ,abstract, journal, author names etc (technical paper). As nice as this tool is, my current experience of it is the clusters can be somewhat hit or miss. Perhaps taking into account citation relationships could improve the accuracy of this tool?
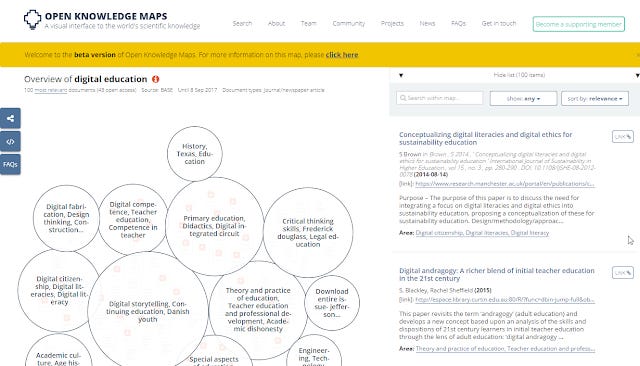
Clustering of papers with keyword digital education from BASE index
3. Citation Gecko (older version reviewed here) - With a nice clean interface this tool which was launched in 2018 is one of my favourite tools. You enter a list of "seed papers" by either searching (data drawn from Crossref) , importing from Zotero libraries or bibtex files and Citation Gecko will map out existing citation relationships between these seed papers. The citation map has two views, allowing you to visualize to find potential interesting papers that frequently cite the seed papers or are cited by the seed papers. You can add these interesting papers to the core seed papers and iterate to expand the citation map.
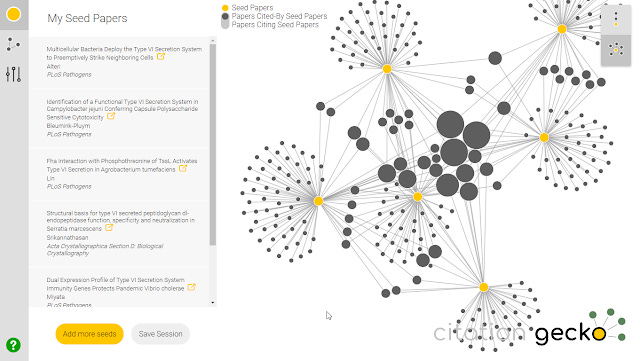
Citation Gecko sample network
The main weakness of this tool for use is firstly you need a fairly good set of papers as seed and more seriously because the current version of Citation Gecko uses Crossref as a source (earliest versions used the far bigger Microsoft academic) and Crossref is known to be missing a high percentage of reference , Citation Gecko tends to produce a lot of nodes(papers) with no citation relationships.
4. Local Citation Network (My brief review, authors' comparison with Citation Gecko) - This tool like Citation Gecko is citation based, however it leverages Microsoft Academic Graph (as well as Crossref) which is more complete than Crossref in terms of metadata and references. Like Citation Gecko you can enter a list of papers (using dois) or even one paper as the input/seed article.
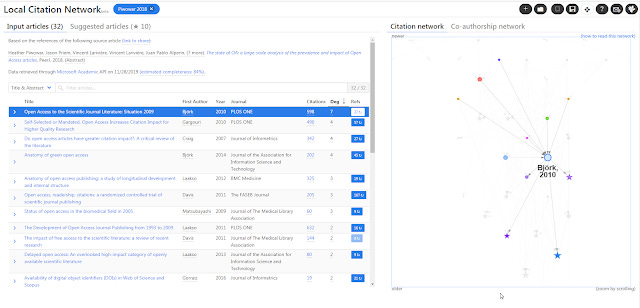
Local Citation Network generated from 10.7717/peerj.4375
It however works only one way. It will look at the reference lists of your input/seed articles and construct a citation network from those papers referenced in your input/seed papers. Suggested papers would be papers that are cited most often locally in this citation network.
As such, the disadvantage compared to Citation Gecko is that it can find suggested papers only older than the source/seed papers while Citation Gecko can find both papers that are cited or cite the seed papers.
It is also interesting to note that Local Citation Network differs from the others already mentioned in that while it can work on a set of seed/source papers , it can also work on just one seed paper, while the rest require you enter a few seed papers (Citation Gecko) or do a keyword search first (Open Knowledge Maps, VOSviewer)
In Local citation network, if you enter just one DOI, it will use the top 10 cited references in the DOI as the seed paper.
The two new tools reviewed in the blog post, Connected Papers and CoCites both work on the same paradigm where you can just enter one article to generate suggestions.
Let's start off with CoCites.
CoCites - using Pubmed to find relevant articles with cocitations
CoCites currently works as a Chrome browser extension that you install which will overlay a "View co-citations" button when you are on PubMed. (Note this overlay will be also seen on Google Scholar)

PubMed search results - with "View co-citation" badges overlaid
Clicking on the "View co-citation" button gets you a page where you can see papers sorted by cocitation frequency. You can then export the results if you login but for me this currently leads to an error page.
In the example below we will be looking at CoCites on the 2015 paper - Autism occurrence by MMR vaccine status among US children with older siblings with and without autism which has PMID: 25898051 as well as DOI of 10.1001/jama.2015.3077
This gets you a list of papers ordered by co-citations. Compared to some of the other tools which can take a while to process, CoCites works almost instantly.
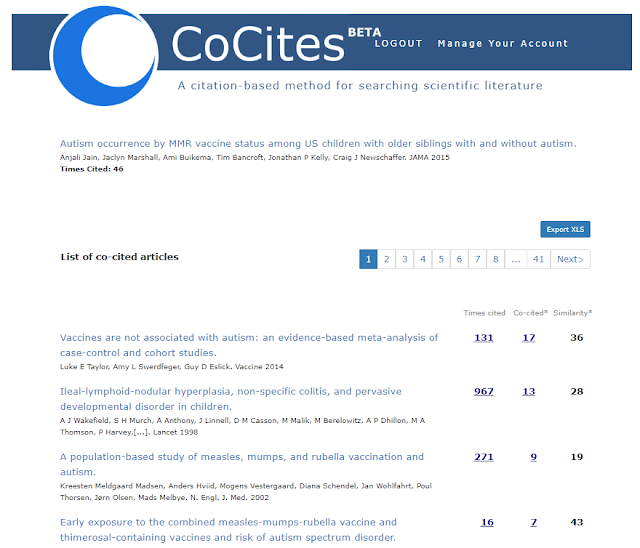
http://cocites.com/coCitedArticles.cfm?pmid=25898051
How is this cocitation calculated?
First note the seed/input/"query paper" you use in this case - Autism Occurrence by MMR Vaccine Status Among US Children With Older Siblings With and Without Autism has 46 citations known to CoCites.
CoCites will look at these 46 cited articles and
a) Extract all the references in these 46 articles (I believe if it is cited more than 100 times, it will use a limit of last 100 cited papers)
b) As I write this CoCites sees about 2,035 unique papers
c) Of these 2,035 unique papers, how many of them are common among the 46 reference lists? In other works, if say Paper A turns up in 23 of these reference lists, it is co-cited 23 times.
The logic is the more frequently a paper appears in the reference lists of the 46 cited papers (or equivalently the more frequently a paper is cocited with the input paper), the more likely it is to be related or similar.
Using the example above, we see the first recommended paper - "Vaccines are not associated with autism: an evidence-based meta-analysis of case-control and cohort studies." is a paper that is cocited 17 times. In other words of the 46 papers that cite the input paper, when you look at the reference list, that paper appears 17 times.
Since the maximum possible number of times a paper can be cocited is 46, if that paper appears 46 times it will have a similarity of 46/46 = 100%. That isn't likely, but in this case the top paper has a similarity of 17/46= 36%
Graphically it looks like this.
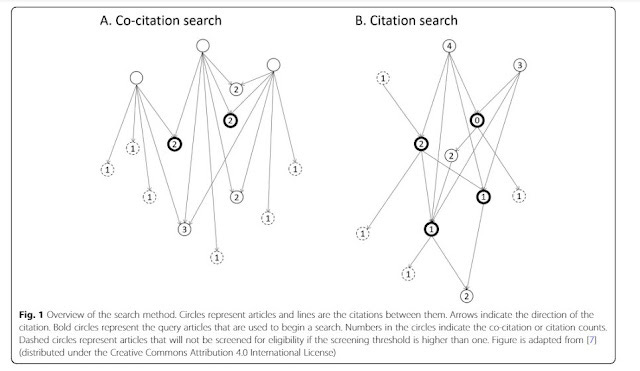
Though this method sounds simple in practice a series of papers have been released studying this method and it has proven quite effective.
For example in Novel citation-based search method for scientific literature: a validation study they tried to "reproduce the literature searches of published systematic reviews and meta-analyses and assess whether CoCites retrieves all eligible articles while screening fewer titles."
They studied over 250 reviews and they found in general when the author screened more than 500 articles a method consisting of doing citation and cocitation searches on the top 2 highest cited articles as input articles generally leads to a more efficient search.
CoCites is still very new at the time of writing, but it seems promising to find similar articles when doing systematic reviews or meta-analysis as it seems to be designed for that purpose.
Co-citations will not work well if the input article has few or even no citations
Still CoCites has some minor shortcomings.
One of the advantages of using citations of the input paper rather than references as Local Citation Network does is that references reflect what one research team felt was relevant at that one moment of time to that paper, while citations produce a more nuanced view of the paper from different angle across different times by different teams. This results in a growing dynamic network generated by citations compared to references.
However the disadvantage is it will not work on papers with no citations.
As it currently stands, as Cocites works only on one seed/input/query article if that article has very few or even no citations it is not going to be effective or even work.
After all if the seed paper has no citations it can't be cocited and the fewer you have the less effective the technique is.
For example at the time of writing, this article on COVID-19 has zero cites tracked by PubMed, so CoCites is unable to work.
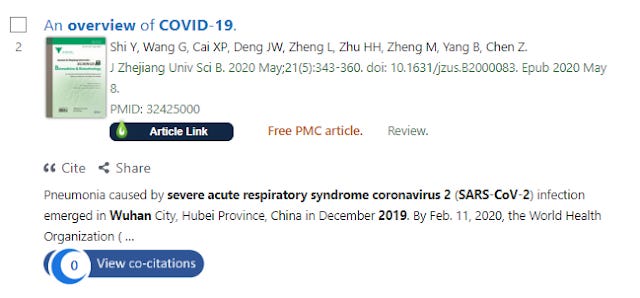
Paper indexed by PubMed with 0 known co-citations
Clicking on the badge about does nothing. (If the "view co-citations" badge is darkened rather than blue it means CoCites has not even indexed the article yet but with the same result).
A workaround of course is just to look at the references of the input paper that is well cited for something else to use. Future versions (see below) might also allow you to combine multiple papers for a CoCites analysis.
Other promised feature improvements coming soon (as of June 2020) include
Perform a co-citation search using multiple query articles.
Find recently published articles through a citation search.
Filter search results using a similarity score
Save search queries for re-use and search updates
Comparison with other tools such as Local Citation Network
One of the more important things when using such tools that draw on citations or even article metadata or full text is to understand the sources it draws from.
In this case CoCites is using PubMed as a source so this makes it unusable for research in areas outside the usual domains covered by PubMed.
The fact that it can work on single papers makes it very close to Local Citation Network. But there is one big difference (besides the fact that one works on Microsoft Academic Graph the other PubMed)
When you enter a single paper into it, Local Citation Network looks at the references of that paper rather than the citations of the paper (which CoCites does) as inputs to extract reference lists. For example let's use the same example as above but entering the doi 10.1001/jama.2015.3077 which is a 2015 paper. into the Local Citation Network.
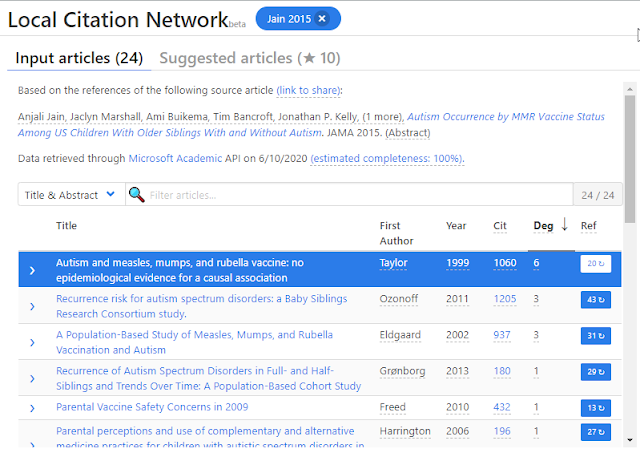
Local Citation Network on Autism occurrence by MMR vaccine status among US children with older siblings with and without autism
Immediately you see the input papers used are all older than 2015 because the system uses references of the 2015 paper and not citations.
Looking at the suggested articles which should be comparable to CoCites we see the following, again you get also older suggestions than the 2015 papers.
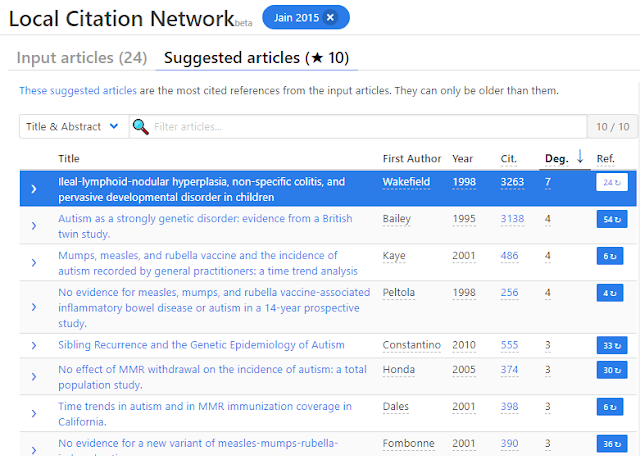
To recap, Local Citation Network used the 24 input articles (which are themselves references of the 2015 paper) to extract references to create a network and within that set of articles it shows suggested articles that are cited the most in that network.
This naturally means that the suggested articles cannot be newer than 2015 (barring errors in the data).
On the other hand using CoCites, while the top cocited papers might and probably also be relatively older papers, it is certainly possible for them to be newer than the 2015 paper inputed.
For example, when using CoCites the 2015 input paper might be cited by a couple of 2020 papers, and almost all of them also cite a 2018 paper, so that 2018 paper would be highly ranked.
This is an advantage of how CoCites works.
On the other hand, as already mentioned CoCites can never get off the ground at all for very new articles with no citations, while Local Citation Network is always able to do something because it uses references (assuming available) of the input article.
So for example while co-cites will not work on this article (as of time of writing as it has zero cites), it works fine in Local Citation Network
Connected Papers - an introduction
In general, many tools use citation networks whether citations of the input paper (CoCites) or references of the input paper (Local Citation Network) to start off the network. As we have seen, this leads to trade-offs in term of which papers the technique can work on. But what if we combined the two? This is where Connected Papers comes in.
Like CoCites and Local Citation Network, connected maps uses one paper and it is stated to
"analyze an order of ~50,000 papers and select the few dozen with the strongest connections to the origin paper." (in my experience you tend to see 40-50 nodes)
But what are the connections used? Citations? References?
Connected Papers seems to try to split the difference to construct a similarity metric
"based on the concepts of Co-citation and Bibliographic Coupling.According to this measure, two papers that have highly overlapping citations and references are presumed to have a higher chance of treating a related subject matter."
and
"builds a Force Directed Graph to distribute the papers in a way that visually clusters similar papers together and pushes less similar papers away from each other."
They state that they are not a citation tree as nodes that do not cite each other directly can still be similar based on this similarity metric and are positioned closely.
To test this out, I tried a new 2020 with no citations.
Here's an example of a graph generated using a relatively new 2020 paper with no citations but 70 references.
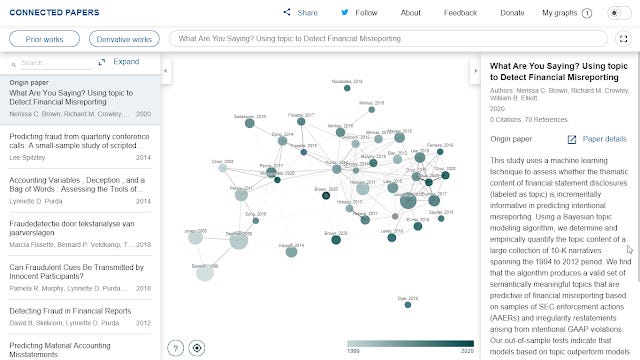
As already mentioned unlike the other tools already mentioned, the edges/lines are not citations but rather based on a similarity metric they calculated (thicker lines are more similar). By clicking on "expand" you can see the exact similarity score.
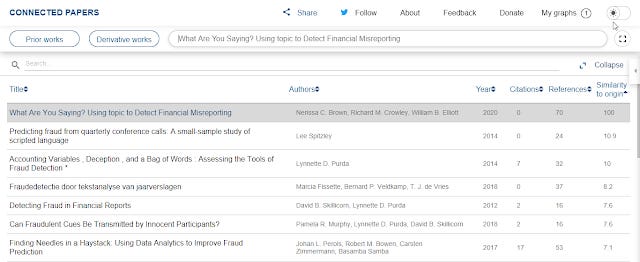
Expanded list view showing "Similarity to origin" metric
The first paper is the original paper so has similarity to origin of 100. The paper closest in similarity to the origin has a score of 10.9.
10.9 as the most similar paper seems to be a little on the low side as most networks I generated with this tool tend to have closest papers with similarity scores that are higher say 15-30%. I guess the lack of citations to this paper is affecting this?
That said eyeballing all the papers in the generated network, most look reasonable to me to be flagged as similar.
The rest of the graph is easier to interpret, the color of the nodes represents year of publication and as you might guess size of the node is the number of citations.

Interestingly, Connected Papers doesn't stop at generating this "dozens" of nodes/papers. You can also additionally find "prior works" or "derivative works"
Prior works are detected by looking at which are the most commonly cited papers in the generated network. Assuming the network generated based on similarity is on target and most similar work cite the most important work, you will in theory get seminal works.
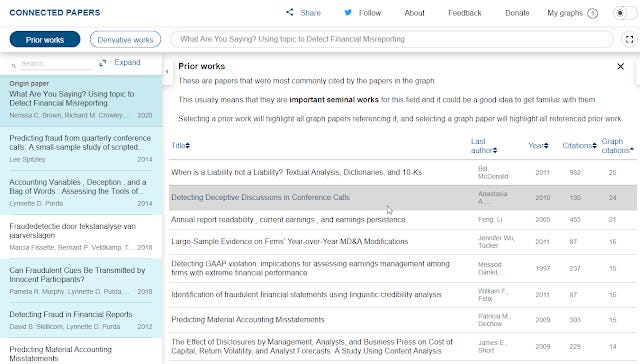
Prior works detected by Connected Papers
If you select a prior work, it will highlight on the left pane , papers from the originally generated network that cited this prior work.
It is also available in the graphical view, where nodes in bright blue were citing the highlighted prior work.
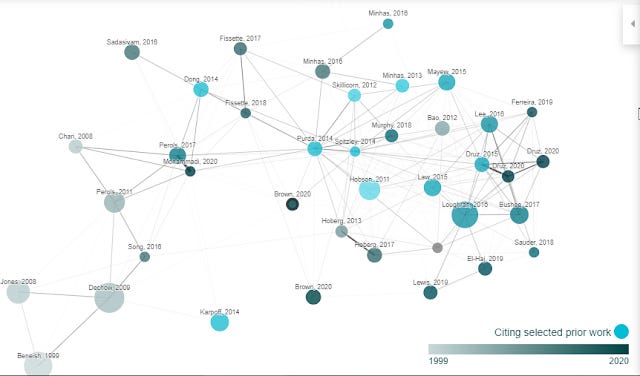
"Derivative work" of course does the opposite, it finds papers that cite the most number of papers in the generated network of papers. It will tend to find reviews or surveys of the field etc. Otherwise it has the exact same functions as "prior works"
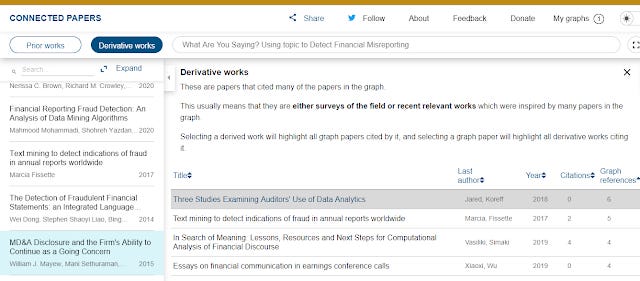
Derivative work generated by Connected Papers
In this example, because we are using a new 2020 paper which itself is on a relatively new development (use of machine learning techniques in accounting research) - there aren't that many review papers out there, yet it still does not do too horribly.
Here's the derivative works found when I input a more normal 2008 paper (on the subject of Wikipedia) of mine.
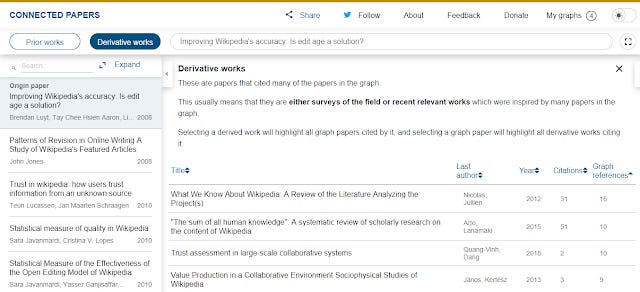
Derivative work of a 2008 paper - Improving Wikipedia's accuracy is Edit age the answer?
While testing so far I almost always get new papers from prior works or derivative works function that are not in the original generated graph. The authors mention to me that it is not impossible for that to happen, or even to have the input paper be in the prior work list but it's rare.
This surprises me a little, as I expected some of the prior work and derivative works to be very similar to the original work, but there is I guess a lot of different ways that a human thinks as "similar" that isn't when seen through the lens of an algorithm that is trying to base similarity based on similarity of reference lists and cocitations.
Comparison of Connected Papers with Cocites
As already mentioned CoCites unlike Local Citation Network and Connected Papers cannot work on articles with no citations. While Connected Papers can do so because it's similarity metric uses both cocitations and bibliometric coupling.
Does this mean CoCites is less powerful? First off I find Connected Papers doesnt always work either on all papers either. I've seen examples where CoCites produces something (granted something not too good) while Connected Papers doesn't.

Secondly like any tool, it can produce not so good results as well. For example, the graph below generated from a paper "Applying Machine learning in accounting research" doesn't give a network that looks high quality (from my view point anyway).
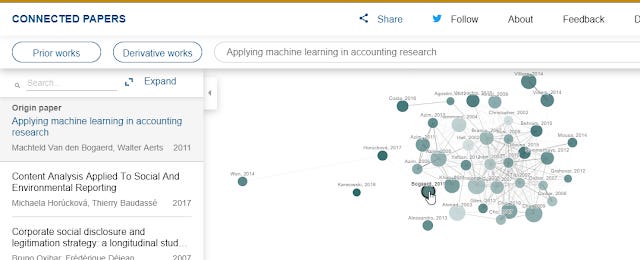
Connected Papers generated network for paper - Applied machine learning in accounting research
We don't exactly know how the similarity metric works but do Cocites and Connected Papers give that different results?
Comparing Connected Papers and CoCites output on the same paper
I tried with the following paper on autism and vaccines - https://pubmed.ncbi.nlm.nih.gov/25898051/
which gave me this
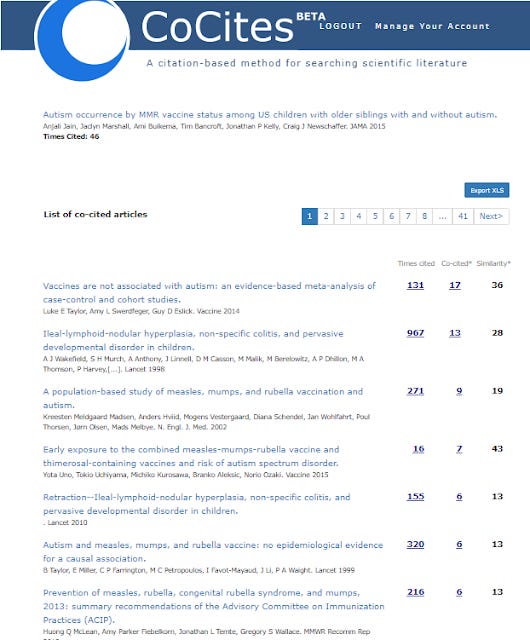
Doing the same for Connected Papers gets me this
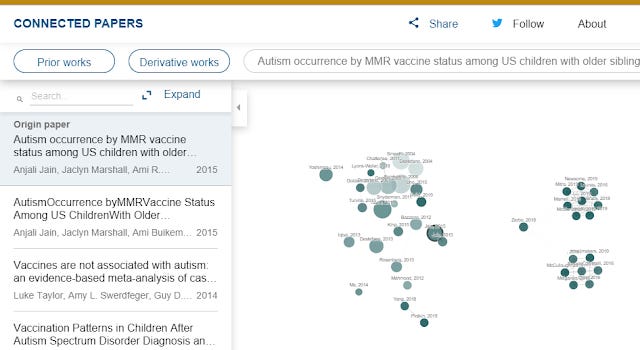
Using the top 10 ranked papers based on cocitations in CoCites, I compared it to the results generated from Connected Papers.
CoCites suggestions look more predictable to me (but I'm not an expert), including listing the famous retracted Wakefield paper linking autism to vaccines in 1998.
Yet papers like that are not listed by Connected Papers in the inital similarity generated network but are in the "prior" works.
This isn't very surprising as I noted earlier that Connected Paper generally doesn't consider influential prior works to be similar (though you can find them using the prior works function).
But what if you include the pool of all the papers Connected Papers considers similar or prior works or derivative?
I found that of top 10 papers considered similar by CoCites, 6 were listed by Connected Papers in either the similarity network or for prior works.
It's hard to say which did better since in this area, there are quite a few works that could fit nicely and I am not familiar with the medical literature sufficiently to say.
Based on this one example we can see that CoCites tends to give slightly older papers while for Connected Papers the similarity network tend to give much newer papers (all of the top 10 similarity to origin papers were after 2010), but if you include the prior works they would overlap somewhat with CoCites.
I speculate based on this and other simple tests that Connected Papers similarity network tends to result in papers that are similar to some extent but they do tend to give you a bigger sampling of topics around the original paper. They often miss out foundational works and even reviews but there is a seperate way to get them, so all is good.
In comparison, as CoCites puts a heavier weighing on citations, it is easier to surface seminal works as well as favouring older papers compared to Connected Papers which uses a blend of co-citations and bibliometric coupling (similarity of reference lists) which does not work against newer papers.
Also given that CoCites as states on their webpage is an ideal method for
"Scientific reviews, including systematic reviews, meta-analyses, and rapid reviews: projects where the goal is to find similar articles"
and
"Finding the best-known or key articles on niche topics"
It probably is designed and works better with a tighter definition of "similar"? I believe Connected Papers is designed and tested more on machine learning topics and it's similarity network gives you a slightly broader definition of what is similar, where you want to sample slightly different types of papers. In any case you can always request specifically for prior works.
All in all both tools are at a very early stage and already look wonderful, and when it comes down to it, if you want to study an area not under PubMed, you have no choice but to use Connected Papers, On the other hand, I would use CoCites for life sciences since it was designed specifically for that (plus the superior metadata quality of PubMed probably counts for something over Semantic Scholar / Microsoft Academic). Both tools currently can't allow you to iterate to grow the network, this is where Citation Gecko shines.
Bonus
As I was writing this blog post, I was informed of yet another tool - PaperGraph. A quick view finds that it uses again Semantic Scholar (similar to Connected Papers).
Like Local Citation Network, CoCites and Connected Papers you enter one paper and it generates a network.
Here is how it does so
"The graph is generated by finding the root node based on the search query, and the following the top 20 citations by total citation count of each paper over two hops (citations of citatations). "
In other words, when you input a paper, it looks at the references (like Local Citation Network) and for 20 of those papers it goes down another layer (again 20) to generate the network.
Below shows how it looks in Hierarchical Layout
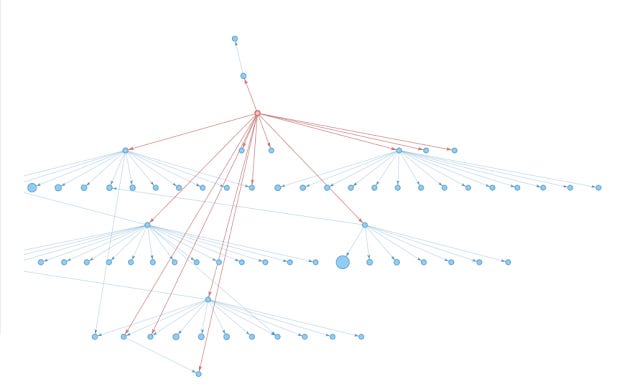
Papergraph network generated from Improving Wikipedia Accuracy: Is edit age the answer? Hierarchical Layout
In the more normal layout, you can look at the list and select "in-degree" to see papers most cited in this network that was generated. It isn't a surprise you get only older papers (than 2018 which is input paper). You can also double click a node to make it the root node (the input paper).
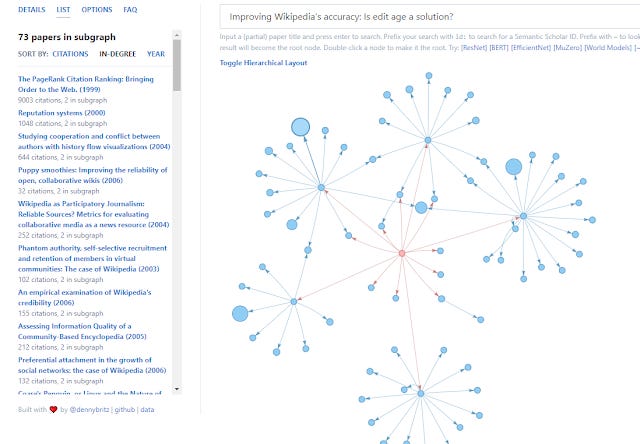
Papergraph network generated from Improving Wikipedia Accuracy: Is edit age the answer?
This seems limiting but by adding the tilde symbol ( ~ ) you can switch the algo to create the network in the other way by citations to the input paper and you then get newer recommendations!
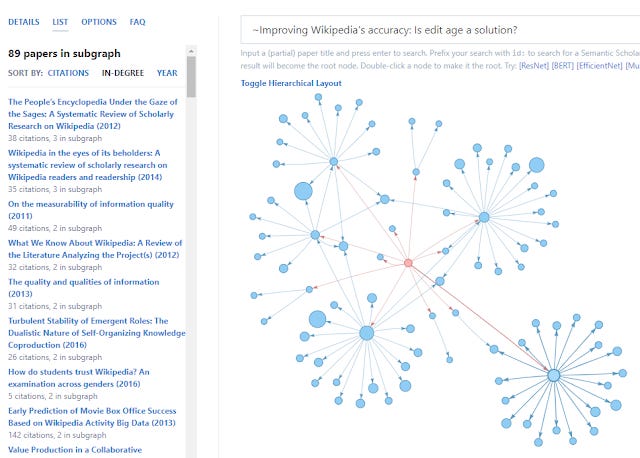
Conclusion
Looking at the comparison table all the way on the top, we can see many of these open tools are powered by data (metadata, citations, full text) from open sources like PubMed, Crossref, Microsoft Academic and Semantic Scholar. This shows the value of having public open data sources that can be used by anyone.
Additional details
Description
It's a cliche to say that the volume of scientific literature produced is increasing at a rate beyond which researchers can keep up. With the rise of Machine learning techniques and more importantly the increasing availability of open meta-data and increased open full text , we are starting to see tools appear that try to leverage this to produce applications that can help take the load off researchers on what to read and to
Identifiers
- UUID
- 561eb3ed-d3f9-4664-a451-89eec81fc1f7
- GUID
- 164998268
- URL
- https://aarontay.substack.com/p/more-researchliterature-mapping-tools_16
Dates
- Issued
-
2020-06-16T13:09:00
- Updated
-
2020-06-16T13:09:00

